CCGP ESTPs
Expressed Sequence Tag Polymorphisms
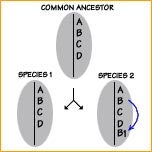 Orthology describes the evolutionary origin of a locus. Loci in two species are said to be orthologous (identical by descent) when they have arisen from the same locus of a common ancestor. For example, gene A in Species 1 and 2 are orthologous, whereas gene B1, which has arisen by gene duplication in species 2, is paralogous to gene B in Species 1. Clearly, comparative mapping in species groups having large genomes with abundant multigene families, such as pines, can be confounded by the mapping of paralogous loci.
Orthology describes the evolutionary origin of a locus. Loci in two species are said to be orthologous (identical by descent) when they have arisen from the same locus of a common ancestor. For example, gene A in Species 1 and 2 are orthologous, whereas gene B1, which has arisen by gene duplication in species 2, is paralogous to gene B in Species 1. Clearly, comparative mapping in species groups having large genomes with abundant multigene families, such as pines, can be confounded by the mapping of paralogous loci.
Along with David E. Harry, Berhanu Temesgen and Garth Brown (Harry et al., 1998; Temesgen et al., 2001; Brown et al., 2001), we have developed a set of primer pairs from loblolly pine expressed sequence tags (ESTs) that can serve as a framework around which comparative genetic maps can be built for pines and other conifers. Polymorphisms in these PCR-based markers are quite easily detected using denaturing gradient gel electrophoresis and other means. By targeting PCR amplification to the 3″ end of genes, the EST polymorphisms (ESTPs) mapped to date appear to be orthologous even though the cDNAs from which the primer sequences were derived represent gene families in loblolly pine. The pedigrees of CCGP collaborators have been screened for successful PCR amplification and polymorphism and the results can be found here.
In addition, a group of cDNAs that are single or low copy in loblolly pine and other pine species have been identified. These markers, listed below and suitable for constructing comparative maps using RFLPs, are freely available upon request.
Loblolly Pine cDNAs Likely to Reveal Orthologous Loci In Other Species
| PtIFG_66 PtIFG_138 PtIFG_606 PtIFG_624 PtIFG_653 PtIFG_669 PtIFG_1454 PtIFG_1584 |
PtIFG_1633 PtIFG_1635 PtIFG_1643 PtIFG_1672 PtIFG_1849 PtIFG_1917 PtIFG_1950 PtIFG_1955 |
PtIFG_1956 PtIFG_2006 PtIFG_2020 PtIFG_2022 PtIFG_2068 PtIFG_2113 PtIFG_2150 PtIFG_2220 |
PtIFG_2253 PtIFG_2274 PtIFG_2291 PtIFG_2358 PtIFG_2442 PtIFG_2540 PtIFG_2547 PtIFG_2564 |
PtIFG_2610 PtIFG_2697 PtIFG_2707 PtIFG_2723 PtIFG_2897 PtIFG_2986 |